In the research article in the Journal of Computational Chemistry entitled “Geometrical benchmarking and analysis of redox potentials of copper(I/II) guanidine-quinoline complexes: Comparison of semi-empirical tight-binding and DFT methods and the challenge of describing the entatic state (part III)” Herres-Pawlis & Leonhardt et al. describe copper model complexes for the entatic state theoretically.
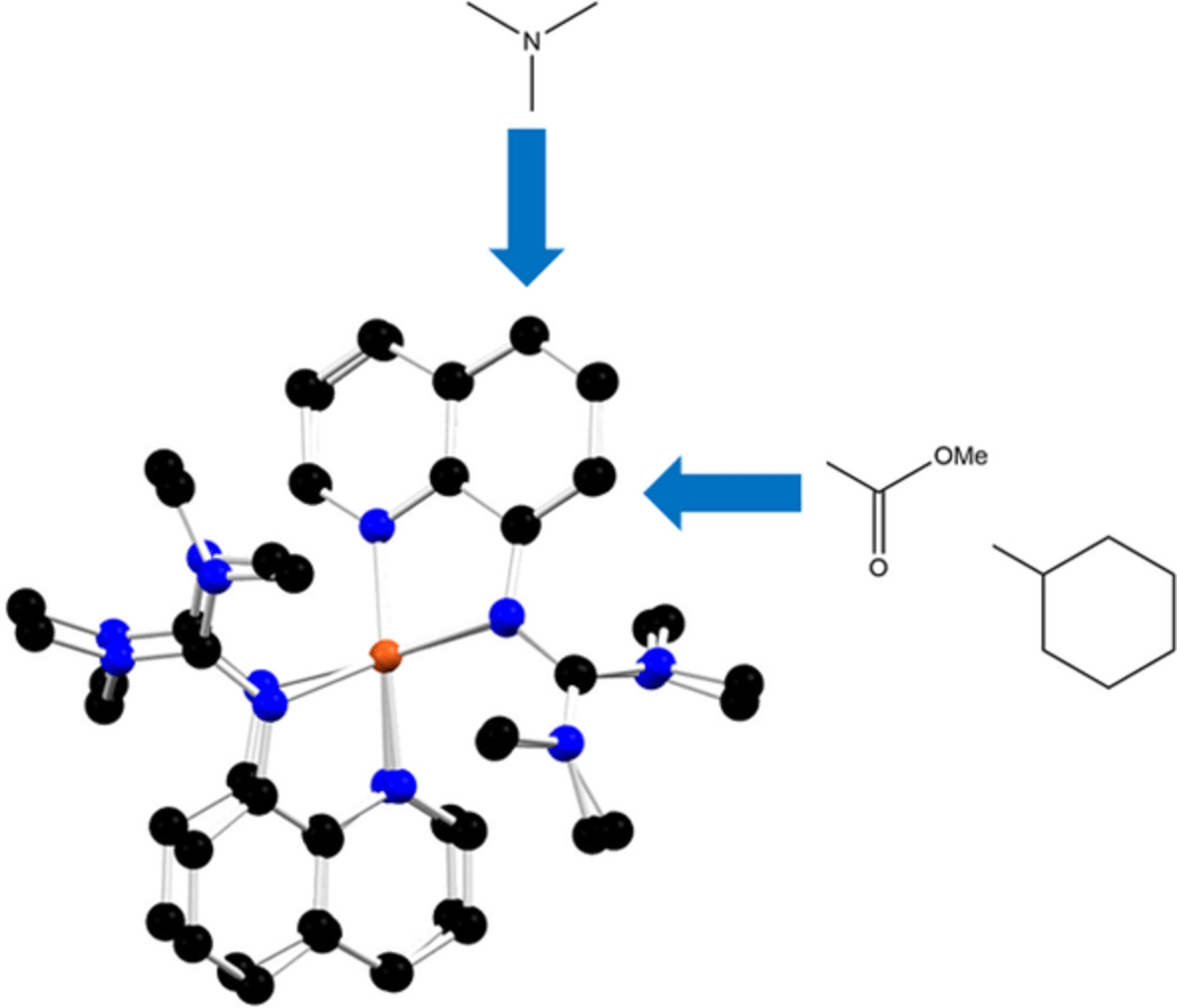
They use the semi-empirical tight-binding method GFN2-xTB, the density functional TPSSh and the double-hybrid functional B2PLYP and evaluate the methods on five different complex pairs (Cu(I) and Cu(II) complexes), and compare how well calculated energies can predict the redox potentials. They find that B2PLYP and TPSSh yield better accordance with the experimental structures but that GFN2-xTB performs surprisingly well in the geometry optimisation at a fraction of the computational cost. TPSSh offers a good compromise between computational cost and accuracy of the redox potential for real-life complexes.
The DFT coordinates have been deposited at the RADAR4Chem repository (dataset). RADAR4Chem accepts every kind of chemical data annotated with their metadata.